Innovative Agricultural Microbiomes
Understanding and manipulating interactions between soil, plant and animal microbiomes to improve water quality and sustainable pastoral productivity
Project Details Ngā taipitopito
Collaborators Ngā haumi
AgResearch | DairyNZ | Lincoln University | Manaaki Whenua Landcare Research | University Of Otago
What are we doing?E aha ana mātou?
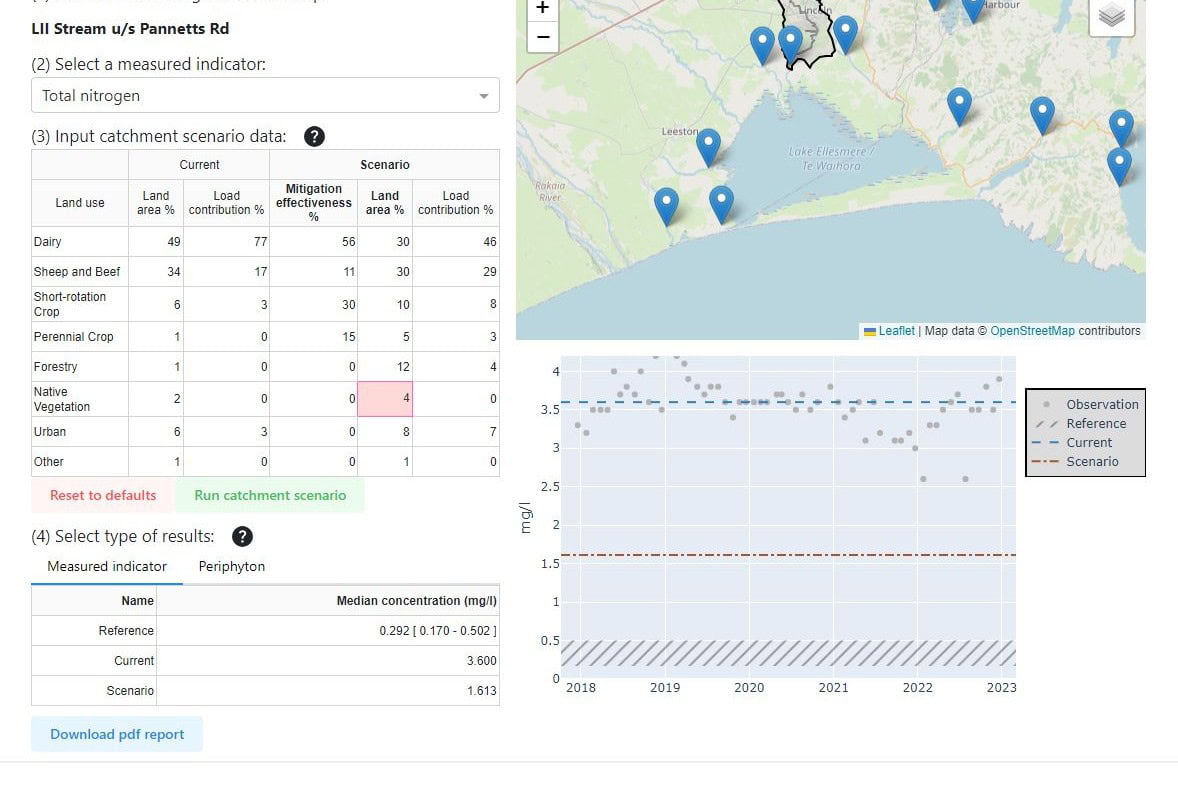
New Zealand dairy farm systems are strongly influenced by various sources of nitrogen: atmospheric nitrogen fixed by legumes, cow effluent, and fertilisers. These inputs must be balanced to maximise plant and animal productivity, and minimise nitrogen leaching to waterways.
The nitrogen cycle relies on the action of microbes: nitrogen-fixing bacteria in the roots of plants, bacteria in the cow rumen, and ammonifying and nitrifying bacteria in the soil. These plant, soil and animal microbiomes are usually viewed as distinct entities, but Innovative Agricultural Microbiomes research is taking the novel approach of investigating them as a meta-microbiome, or agricultural microbiome.
Our researchers are learning how agricultural microbiomes respond to increasing levels of nitrogenous fertiliser, and are identifying the soil-plant-animal microbiome interactions that mediate nitrogen in a dairy production system.
Our research aims to manipulate microbiome-driven processes in the soil, plant and animal to minimise the leaching of nitrogen, provide nutrients for forage plants, and supply the micronutrients and energy that animals need to produce foods consumed by humans.
How can the research be used? Ka pēhea e whai take ai te rangahau?
- An animal trial has been completed in a ryegrass-white clover dairy production system, utilising the dairy farm facilities at the Lincoln University Ashley Dene Research and Development Station (ADRDS). The work involves the characterisation of soil, plant and animal microbiomes under an external nitrogen (in the form of urea) treatment regime.
- DNA and RNA from the soil, plant and animal microbiomes will be sequenced and correlated with metabolites, nitrogen leaching, pasture production and composition, and milk production data. We are identifying key microbial populations, metabolic pathways and genes associated with both enhanced production and reduced environmental impacts.
- An assessment of microbiomes and their interactions between the animal, soil and plant environments will show how microbiomes align with the Māori concept of mauri (life force).
- The microbiome marker genes that are correlated with productivity and/or sustainable practices will be useful as supporting evidence for product verification schemes to show that NZ’s pastoral farming systems are natural and sustainable.
- Our research will support New Zealand’s competitive advantage in researching novel endophytic microorganism for existing and future crop species. The potential future value of novel endophytic species in assisting plants to adapt to global temperature rises and increased pest pressures will be significant. (For example, the current value to New Zealand of commercialised ryegrass endophytes AR1 and AR37 is estimated to be $155–$285 million annually.)
 View Our Strategy Document 2019 – 2024
View Our Strategy Document 2019 – 2024